Abstract base class for Differential Evolution mutation strategies. More...
#include <Mutation.h>
Public Member Functions | |
| virtual | ~Mutation ()=default |
| Virtual destructor for safe polymorphic deletion. | |
| virtual std::vector< double > | mutate (const std::vector< std::vector< double > > &population, size_t targetIndex, double F, const std::vector< double > &bestVector, MersenneTwister &mt, double lowerBound, double upperBound)=0 |
| Generates a mutant vector using the mutation strategy. | |
Protected Member Functions | |
Shared Utilities | |
| std::vector< size_t > | getSubset (size_t populationSize, size_t subsetSize, size_t source, MersenneTwister &mt) |
| Selects a random subset of population indices, excluding a source index. | |
Detailed Description
Abstract base class for Differential Evolution mutation strategies.
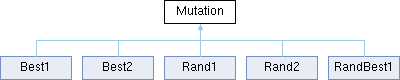
Mutation defines the interface and shared utilities for all DE mutation operators. Each concrete mutation strategy (Rand1, Best1, Rand2, Best2, RandBest1) implements the mutate() method to generate a mutant vector from a population based on a selected mutation formula.
Mutation Role in DE:
In Differential Evolution, mutation is the primary source of variation:
- Select random individuals from the population
- Apply a weighted combination of differences to create new variation
- The mutation formula determines which individuals and differences are used
Mutation Formula Notation:
Standard DE mutation formulas follow the pattern: DE/base/N, where:
- base: The base vector used as starting point (either random or best)
- N: The number of difference vectors applied (1 or 2)
Common formulas:
- DE/rand/1: mutant = r0 + F*(r1 - r2)
- DE/rand/2: mutant = r0 + F*(r1 - r2) + F*(r3 - r4)
- DE/best/1: mutant = best + F*(r1 - r2)
- DE/best/2: mutant = best + F*(r1 - r2) + F*(r3 - r4)
- DE/rand-to-best/1: mutant = x + λ*(best - x) + F*(r1 - r2)
Scaling Factor F:
The differential weight F (typically in range 0.0 to 2.0) controls the amplitude of variation. Key considerations:
- F ≈ 0.5-0.8: Conservative exploration, good for fine-tuning
- F ≈ 0.8-1.5: Moderate exploration-exploitation balance
- F > 1.5: Aggressive exploration, may miss good regions
Individual Selection:
The getSubset() helper selects random distinct individuals from the population (excluding the target individual). This ensures the mutant is derived from different genetic material than the target.
Member Function Documentation
◆ getSubset()
|
inlineprotected |
Selects a random subset of population indices, excluding a source index.
This is a shared utility used by all mutation strategies to randomly select distinct individuals from the population. The selection uses a partial Fisher-Yates shuffle for efficiency and guarantees no duplicates.
Algorithm:
- Create a list of all valid indices [0, populationSize-1] except source
- Perform first subsetSize iterations of Fisher-Yates shuffle:
- For each position i, randomly swap with position j in range [i, size)
- Return the first subsetSize elements (which are now randomized)
Complexity:
Time: O(populationSize + subsetSize) for setup and shuffle Space: O(populationSize) for temporary index vector
Parameters:
- Parameters
-
[in] populationSize Total number of individuals in the population. Range: subsetSize + 1 to infinity. [in] subsetSize Number of distinct indices to select. Range: [1, populationSize - 1]. Must be less than populationSize (to exclude source). [in] source Index to exclude from the selection. This individual is not included in the returned subset. Range: [0, populationSize - 1]. [in,out] mt Mersenne Twister random number generator. Passed by reference; state is modified. Used to generate random swap positions.
Return Value:
- Returns
- A vector of subsetSize distinct indices from [0, populationSize-1], excluding source. All elements are unique. Size: exactly subsetSize.
Guarantees:
- All returned indices are distinct (no duplicates)
- No returned index equals source
- All indices are in range [0, populationSize - 1]
- The selection is uniformly random (all subsets equally likely)
Usage in Mutation Strategies:
Mutation formulas require 2-5 distinct random individuals:
Example:
Preconditions:
- populationSize >= subsetSize + 1 (at least room for subset + source)
- source < populationSize
- subsetSize > 0
Implementation Note:
The partial Fisher-Yates algorithm is used instead of full shuffle or repeated sampling for O(populationSize + subsetSize) complexity without the risk of duplicate rejection sampling.
◆ mutate()
|
pure virtual |
Generates a mutant vector using the mutation strategy.
Pure virtual method that each concrete mutation strategy implements to produce a mutant vector. The mutant combines selected population individuals according to the strategy's formula.
Algorithm Pattern:
All mutation strategies follow this general workflow:
- Select N random distinct individuals from the population (via getSubset())
- Compute a weighted combination of these individuals
- Clamp the result to the domain bounds
- Return the mutant vector
Parameters:
- Parameters
-
[in] population The complete population of candidate solutions. Each element is a vector of dimension equal to the optimization problem. Size: popSize × dimension. [in] targetIndex Index of the target individual in the population. This individual is excluded from random selection (cannot use its own genetic material for mutation). Range: [0, popSize-1]. [in] F Differential weight (mutation scaling factor). Controls the amplitude of variation in the mutation formula. Typical range: (0, 2], common values: 0.5-1.0. - Larger F: more aggressive exploration
- Smaller F: more conservative fine-tuning
[in] bestVector The best (lowest fitness) individual found so far. Used in "best" strategies (Best1, Best2, RandBest1). For "rand" strategies, may be ignored. Dimension: same as population elements. [in,out] mt Mersenne Twister random number generator. Used to randomly select individuals and generate variations. Passed by reference; state is modified during execution. [in] lowerBound Lower bound of the feasible domain. All dimensions share this uniform bound. Used to clamp mutant values to valid range. [in] upperBound Upper bound of the feasible domain. All dimensions share this uniform bound. Used to clamp mutant values to valid range.
Return Value:
- Returns
- A new mutant vector with dimension matching the population. All values are within [lowerBound, upperBound] due to clamping. Size: equal to population[0].size().
Postconditions:
The returned mutant vector satisfies:
- Dimension matches population dimension
- All elements are within [lowerBound, upperBound]
- Is derived from 2-5 random population members (depends on strategy)
Strategy-Specific Behavior:
Different concrete strategies implement different formulas:
- Rand1: Uses 3 random individuals; more exploratory
- Best1: Uses best individual as base; more exploitative
- Rand2: Uses 5 random individuals; more variation, slower
- Best2: Uses best with 4 additional individuals; balanced
- RandBest1: Interpolates between best and random; hybrid approach
Complexity:
Time: O(dimension) - linear in the problem dimension Space: O(dimension) - for the returned mutant vector
Usage Example:
- See also
- getSubset() for the random selection mechanism
The documentation for this class was generated from the following file:
- src/opti_py/cpp/include/opti_py/Optimizer/DifferentialEvolution/Mutation/Mutation.h